SCoPE2:
Single Cell ProtEomics by Mass Spectrometry (Version 2)
Quantifying proteins in single cells at high-throughput with mass spectrometry
SCoPE2 Preprint SCoPE2 Paper SCoPE2 code on GitHub
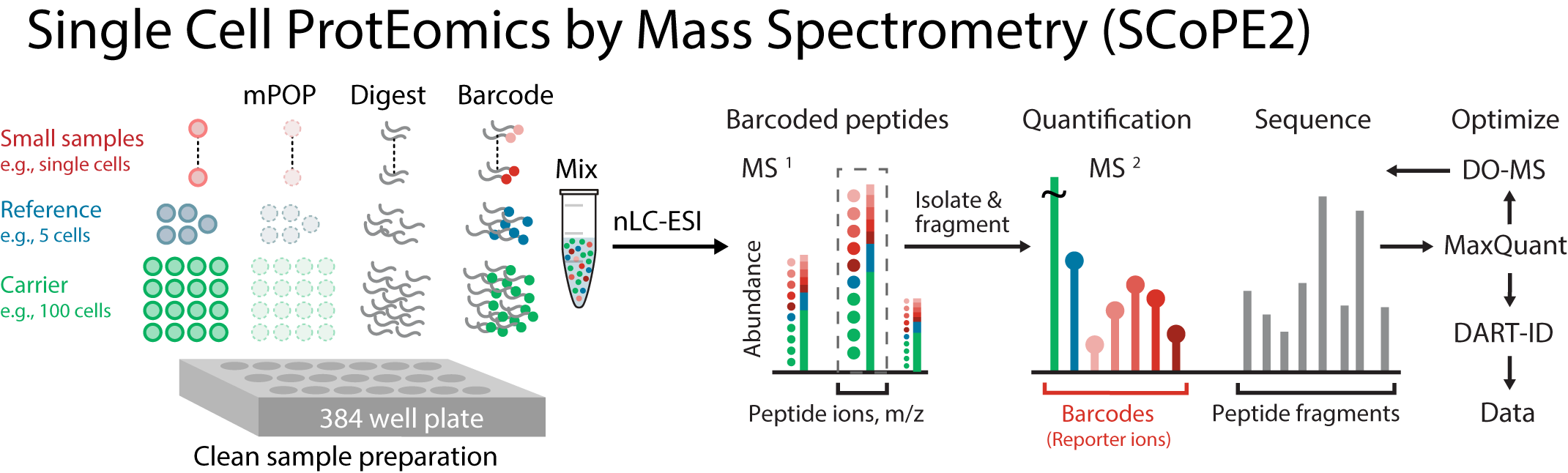
The fate and physiology of individual cells are controlled by proteins. Yet, our ability to quantitatively analyze proteins in single cells has remained limited. To overcome this barrier, we developed SCoPE2. It substantially increases quantitative accuracy and throughput while lowering cost and hands-on time by introducing automated and miniaturized sample preparation. These advances enabled us to analyze the emergence of cellular heterogeneity as homogeneous monocytes differentiated into macrophage-like cells in the absence of polarizing cytokines. SCoPE2 quantified over 3,042 proteins in 1,490 single monocytes and macrophages in ten days of instrument time, and the quantified proteins allowed us to discern single cells by cell type. Furthermore, the data uncovered a continuous gradient of proteome states for the macrophage-like cells, suggesting that macrophage heterogeneity may emerge even in the absence of polarizing cytokines. Parallel measurements of transcripts by 10x Genomics scRNA-seq suggest that our measurements sampled 20-fold more protein copies than RNA copies per gene, and thus SCoPE2 supports quantification with improved count statistics. Joint analysis of the data illustrates how variability across single cells can reveal transcriptional and post-transcriptional gene regulation. Our methodology lays the foundation for automated and quantitative single-cell analysis of proteins by mass-spectrometry.
Perspectives
- Learning from natural variation across the proteomes of single cells, PLOS Biology
- Scaling Up Single-Cell Proteomics, Molecular & Cellular Proteomics
- Driving Single Cell Proteomics Forward with Innovation, Journal of Proteome Research
- Unpicking the proteome in single cells, Science
- Transformative Opportunities for Single-Cell Proteomics, Journal of Proteome Research
Reviews
- Single-cell protein analysis by mass spectrometry, Current Opinion in Chemical Biology
- Single cell protein analysis for systems biology, Essays in biochemistry
Highlights
- Highlight by C&EN
- Highlight by Nature Methods
- Technology feature at Nature Methods
- Single-cell proteomics for the Human Cell Atlas
- News in Proteomics Research
- The single cell proteomics revolution
About the project
SCoPE2 is a project developed in the Slavov Laboratory and SCP Center at Northeastern University, and was authored by Harrison Specht, Ed Emmott, Aleksandra Petelski, R. Gray Huffman, David H Perlman, Marco Serra, Peter Kharchenko, Toni Koller, and Nikolai Slavov.
Contact the authors by email: nslavov{at}northeastern.edu.
This project was supported by funding from the NIH Director’s Award and by an Allen Distinguished Investigator Award from the Paul G. Allen Frontiers Group.